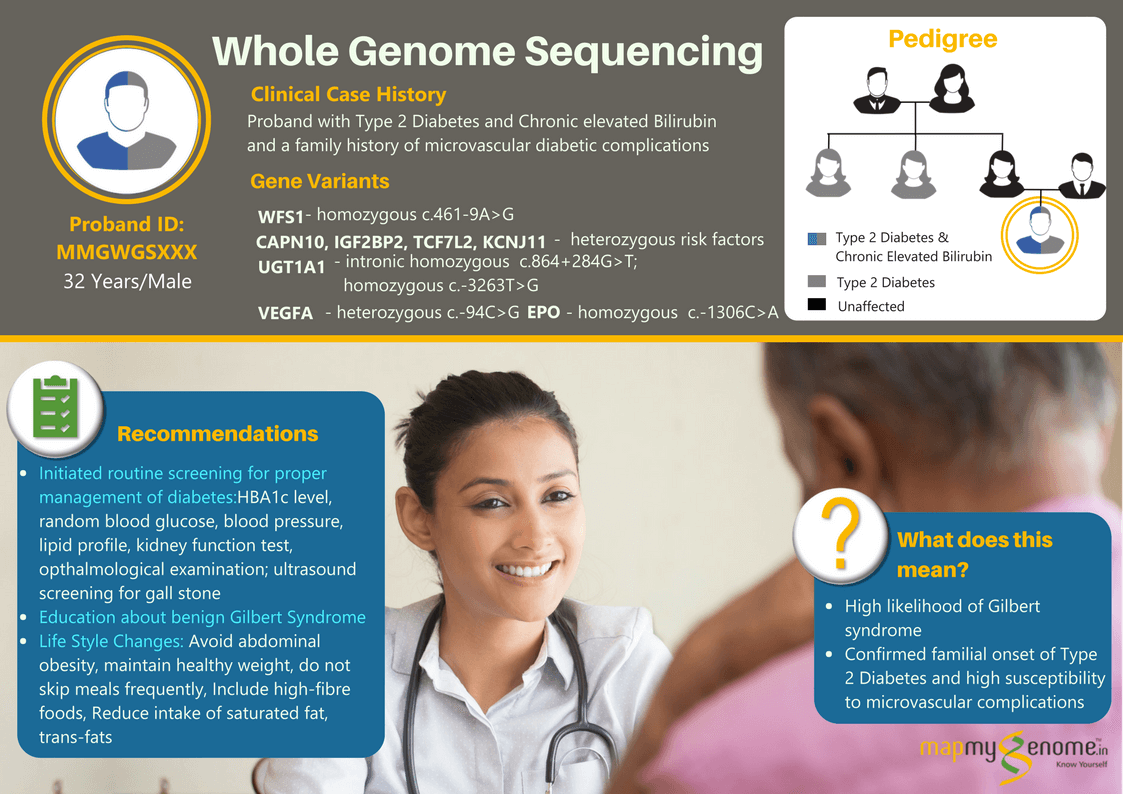
A 32-year-old male affected with type 2 diabetes and chronic hyperbilirubinemia (8 to 10-fold elevated bilirubin levels in the blood) wanted to find out if these could be caused by genetic factors in him? When he went to meet his physician for gaining some insight into the genetics of these conditions, the physician referred him to Mapmygenome for genetic counseling that would help him understand the heritability of these medical conditions and suggest the right genetic test keeping in view his personal medical and family history. During his discussions with a genetic counselor, he revealed that two of his maternal aunts and his paternal grandfather were also affected with Type 2 Diabetes and associated microvascular complications (kidney and eye-related). However, there was no family history of diagnosis of hyperbilirubinemia.
Type 2 diabetes (the more common type) is known to have a stronger genetic susceptibility and link to family history than type 1 diabetes, although both the conditions also depend on diet, lifestyle and environmental factors. If one has a family history of type 2 diabetes, it may sometimes be difficult to figure out whether diabetes in this individual is due to lifestyle factors or inherited susceptibility. In most people, it is due to both. Families tend to have similar eating and exercise habits due to which obesity could also be prevalent in multiple family members, an important risk factor for diabetes.
Adults or adolescents with Hyperbilirubinemia (also commonly called as Gilbert syndrome) can have episodes of elevated levels bilirubin in the blood. These episodes are generally mild and may occur when the body is under stress (example, dehydration, fasting, illness, vigorous exercise). Rarely, people with chronic hyperbilirubinemia may also experience abdominal discomfort or tiredness.
A whole genome sequencing test was advised keeping in mind that both hyperbilirubinemia and diabetes have several different associated genes and cascade testing of each single gene is not recommended as the testing can take a very long time and is also not a cost-effective approach in this age of advanced next generation sequencing (NGS) technology. He agreed and opted for the Whole Genome Sequencing genetic test which was able to simultaneously analyze all known associated genes that may have increased his susceptibility for any of these conditions.
The whole genome sequencing test identified a homozygous ‘pathogenic variant’ in WFS1 gene that is known to increase the risk of Diabetes Mellitus, type 2 by an odds ratio of 1.43. In addition, the test also identified multiple genetic ‘risk factors’ in the UGT1A1 gene associated with increased risk of elevated serum bilirubin levels/ Gilbert syndrome, and in CAPN10, IGF2BP2, TCF7L2 and KCNJ11 associated with increased risk of diabetes mellitus/ insulin resistance and variants in VEGFA and EPO that increase the risk of diabetes-related microvascular complications.
During the post test genetic counseling session, the individual was counseled that these are only risk factors, and have a moderate additive effect on the overall incidence of the specific disease. Having a risk factor/ susceptibility gene variant does not mean that the disorder WILL occur. Whole Genome Sequencing was of immense help to the patient, and his treating physician in the case in question; some of the significant ones are listed below:
1. The referring Doctor gained insight into the reasons for his patient’s chronic hyperhomocysteinemia and was glad to know that it wasn’t a serious condition, requiring no intervention.
2. The client’s general physician felt the need to refer him to an endocrinologist, in view of his moderately elevated genetic risk of diabetes-related complications which was supported by his positive family history; the endocrinologist initiated the patient on routine surveillance for kidney and eye-related complications with the intention of early detection and treatment. The endocrinologist also reviewed the patient’s current diet and lifestyle and saw this as an opportunity to discuss in detail, the do’s and don’ts considering his strong genetic risk variants in addition to his family history of type 2 Diabetes.
3. The individual was made aware of the importance of routine screening tests to enable better management of the condition and specific recommendations were made for dietary and lifestyle changes aimed at reducing disease progression or disease-associated complications in the future.
A) A comprehensive pharmacogenomic section for drugs with relevant literature and FDA recommendations: In this case, the report offered useful information on response to Glimepiride, Glipizide, Chlorpropamide, Glyburide drugs used to treat Diabetes. The referring Doctor found this information extremely useful to be cautious of adverse drug reactions and toxicity, titrate the drug dosage or recommend an alternate drug in view of genetic risk factors for increased or decreased drug sensitivity for these drugs.
B) An extended carrier screening section for common autosomal recessive disorders: The individual tested was detected to be a carrier of Congenital adrenal hyperplasia (CAH) and was counseled of the importance of screening his partner’s carrier status.



